Fifteen years ago, a class of proteins was discovered, which give humans innate immunity to HIV-1. Unfortunately, HIV-1 is a smart virus and has evolved to battle these proteins. Northeastern researchers, with help from their collaborators, have been studying these proteins for several years to help further understand their function and mechanisms in the hopes to be better prepared against HIV-1.
The research, published in Nature Communications, highlights the work of Northeastern Physics Professor and Chair Mark Williams, postdoctoral researcher Mike Morse, Professor Linda Chelico, University of Saskatchewan in Canada, and Ioulia Rouzina, Ohio State University. The paper examines on the capabilities of a human protein that inhibits HIV-1, APOBEC3G (A3G).
Their findings determined that when the A3G protein grows from a single protein (monomer) to a two-protein complex (dimer), its function is transformed from being an editing protein that mutates viral DNA to acting as a roadblock for further replication of the virus.
“I was completely surprised by the result that dimers were sufficient to do this,” said Williams. “We got our data, and the data told us this, but it took us a long time to realize that the only way to understand this data is that dimers must be the model for the protein’s properties to suddenly change.”
The Williams Laboratory for Single Molecule Biophysics specializes in instruments called optical tweezers that study molecules such as single DNA molecules or DNA-protein complexes. This tool holds DNA or RNA between two polystyrene beads to look at interactions with the DNA by observing changes in its length and tension. Using this technique, the lab studies many different biological systems including HIV-1 replication. A combination of these biophysical methods with the enzyme activity experiments and creation of mutant forms of A3G by the Chelico lab allowed for Williams and Morse to compare the original form of the protein with different mutants containing structural changes. This allowed for the isolation of the process that resulted in stably bound dimerized protein.
For the past 12 years, the Williams lab has been studying HIV-1 replication with help from an NIH grant. Their most recent project involves understanding the innate immune proteins that give humans immunity to HIV-1. There are seven proteins in the APOBEC3 family, some of which battle retroviruses like HIV-1, while others battle retrotransposons, which are genetic elements that amplify themselves within a genome and have potential to cause disease.
All APOBEC proteins are cytidine deaminases, allowing them to modify single-stranded DNA to replace bases, causing mutations when the strand is replicated. However, A3G also has another function that does quite the opposite. Rather than quickly zipping along the genome making edits, it can become a stable binding protein that inhibits the reverse transcription process from occurring, preventing HIV-1 replication. This happens because of a process called oligomerization, where single protein units (monomers) join together to create multi-protein complexes or oligomers.
“The idea that you could be zooming along and making these changes, and yet also be blocking something else from moving, didn’t really make sense,” said Williams. “That’s why the oligomerization is so important. As they start off fast and move along, over time they oligomerize and become very slow.”
This team of researchers discovered the oligomerization as published in their 2014 paper in Nature Chemistry. Their study showed that the growth of a multi-protein complex caused the protein to slow down, but it was not known how many proteins were required to make a slow complex. If the number was high, the results might not be relevant for viral replication. In addition, they could show that the complexes were slow, but they could not tell if the slow complexes had enzyme activity. But now their new research has found the answers to these two critical questions.
“Between our experiments where we directly observed the binding of proteins onto DNA, and Professor Chelico’s work looking into the enzymatic activity of A3G, we found that on short time scales in the monomeric form, the protein binds and dissociates very rapidly from DNA and has high enzymatic activity,” said Morse. “But once you let it form these oligomers, the binding is much more stable, and doesn’t dissociate as readily. Professor Chelico was able to find that the enzymatic activity actually decreases when this process occurs so you have these two functions – the enzymatic activity which occurs on one-time scale, and in one form, and this secondary function that, due to the oligomerization, occurs on another timescale.”
Their research showed that this happens even if just two proteins come together, forming a dimer. The dimer is enough to make the protein complex stable and alter the protein’s function. Since only a dimer is needed, multiple roadblocks could be acting to prevent reverse transcription from occurring. Now, this important function is more likely to be biologically relevant.
“Knowing how something inhibits HIV-1 could be a guide to helping design new methods for anti-HIV drugs. The HIV-1 virus has developed its own protein to trick the cell into degrading APOBEC proteins, so if we can figure out how to stop that degradation, maybe we’d be able to enhance the ability of this protein to inhibit HIV-1,” Williams said.
The Williams Lab at Northeastern University plans on continuing to study the different APOBEC proteins, others of which inhibit HIV-1 or retrotransposons. Through studying these proteins with their unique biophysical measurement tools, their lab hopes to understand the proteins’ regulation, activity, and processes to help us to be further prepared against HIV and other related diseases.
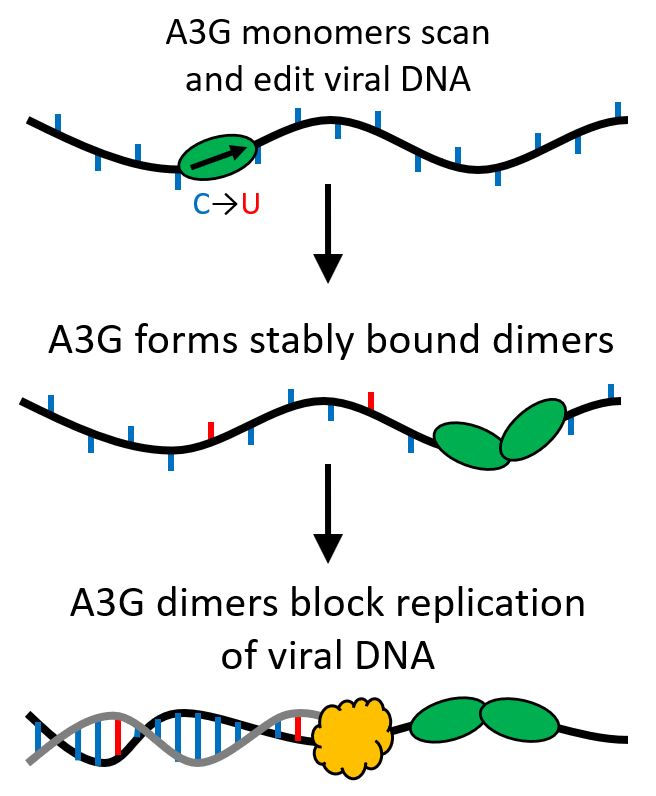
Illustrated representation of APOBEC3G in its first and second functions. Created by Mike Morse.

