
Institutes, Centers, and Consortia
The big questions of today require more than the sum total of our knowledge and ingenuity—they demand data, and lots of it. By harnessing powerful tools and algorithms, researchers can decode scientific complexities faster and at greater scales than ever before. A new, data-driven frontier has emerged, empowering scientists to transform our understanding of the world.
Institutes
-

The Barnett Institute is a center of excellence in the development and application of technologies for biopharmaceutical characterization and proteomics and systems biology. Through its faculty, research scientists, and students, the Institute is improving possibilities every day for how we can address the challenging diseases and complex healthcare issues of the future.
-
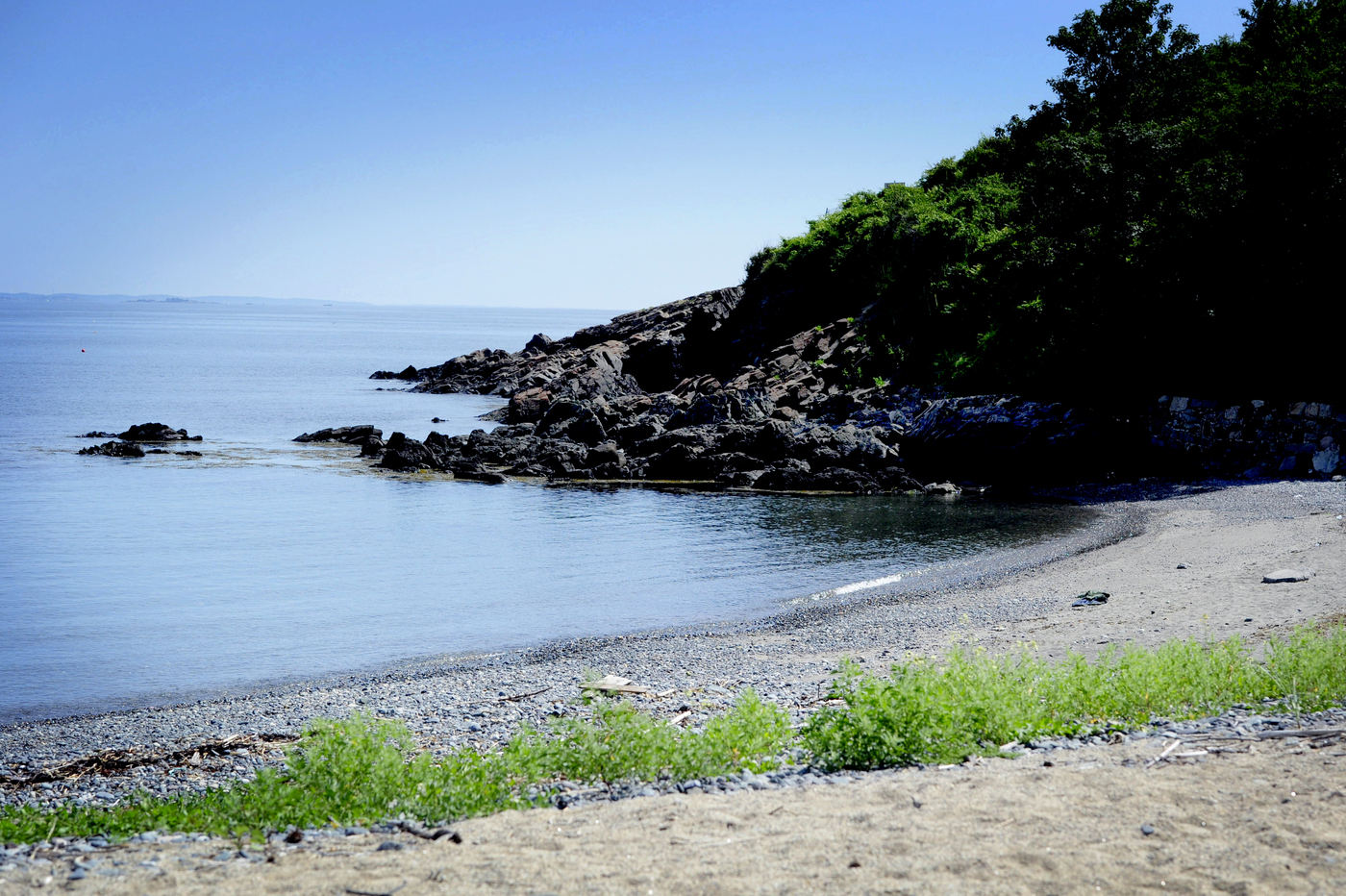

The Coastal Sustainability Institute focuses on creating cleaner, safer, smarter coastal communities. We are committed to uncovering new ways to adapt to and mitigate threats at the land-sea interface, including sea-level rise and storm surge, collapsing fisheries, and coastal pollution.
-
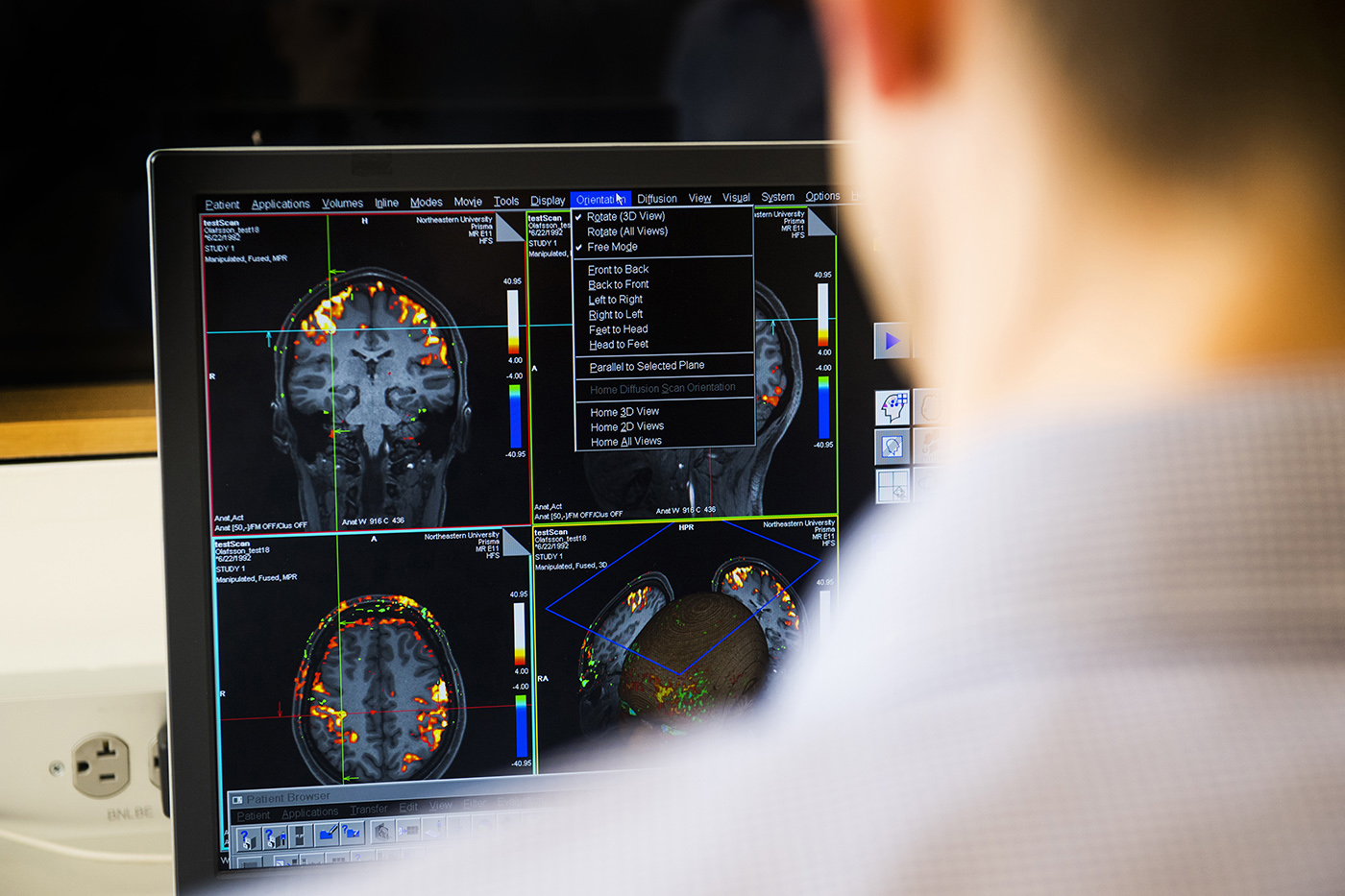
The institute investigates the effects of lifestyle choices and health behaviors (e.g., physical activity, diet) and their physiological sequelae (e.g., fitness, adiposity) on brain and cognition. From a neuroimaging perspective, the researchers’ interests lie in understanding how health influences brain and behavior as it relates to increased health and effective functioning for individuals across the lifespan.
-
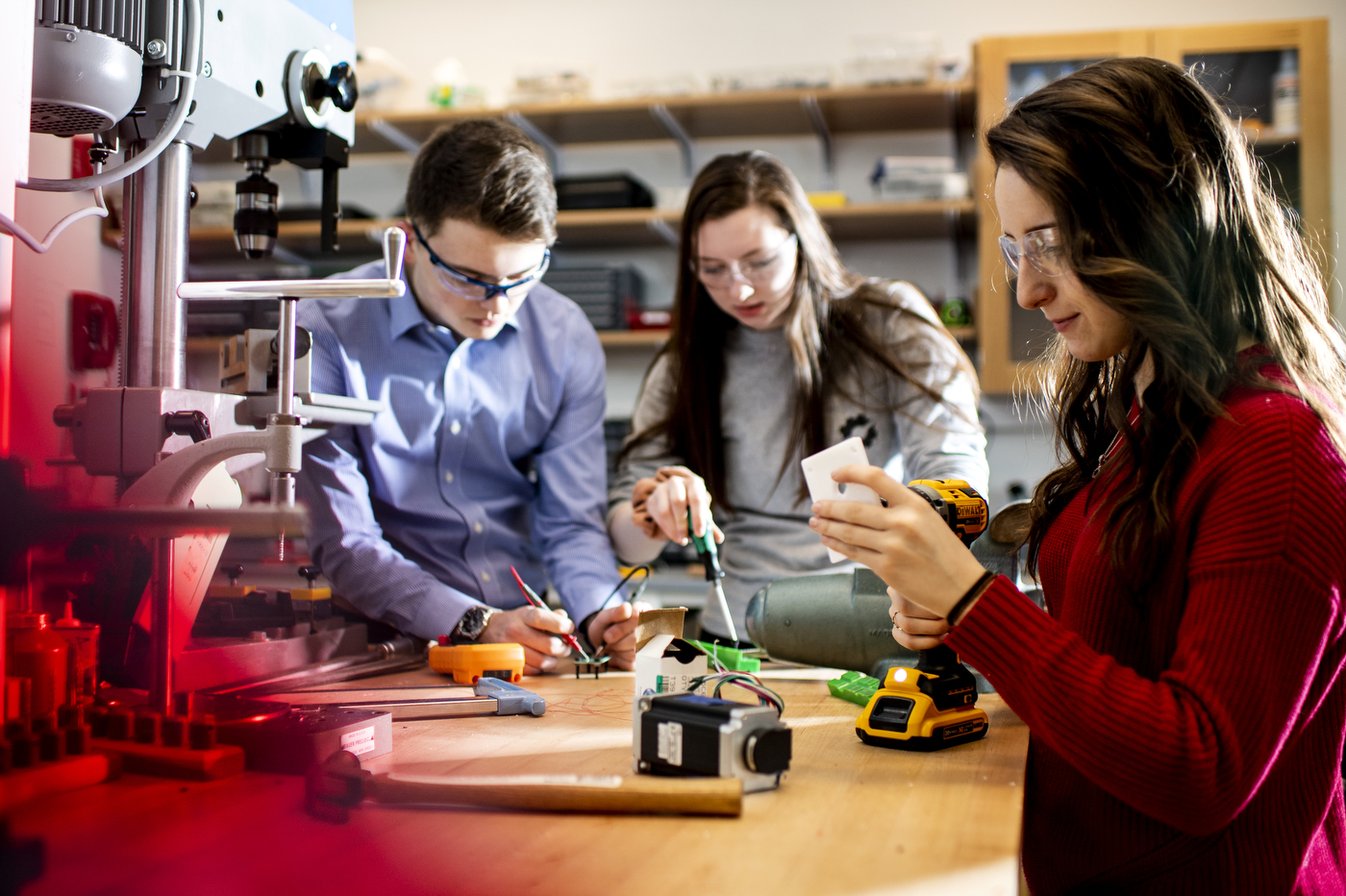

The Institute for Mechanobiology accelerates mechanobiology discovery and technology to advance human medicine and health. Their research investigates the role of force and mechanics in biological systems, discovers the root causes of mechanobiological pathologies that negatively affect quality of life, designs interventions and sensors to alter mechanical inputs and biological outputs, and pursues mechanotherapeutics that restore function or slow the progression of diseases.
-

The mission of the Institute for Plant-Human Interface (IPHI) is to deepen our understanding of plant biology and the plant-human interactions that significantly influence human health and sustainability on Earth. IPHI is committed to devising sustainable strategies that benefit both plants and humans, and to fostering the development of future scientists and engineers in this critical field.
-

The Network Science Institute works to discover and inspire fundamentally new ways to measure, model, predict, and visualize meaningful interactions and interconnectivity of social, physical, and technological systems.
-

The Quantum Materials and Sensing Institute (QMSI) seeks to accelerate practical quantum materials and technologies from concept to commercialization. QMSI researchers reflect expertise in theory, materials synthesis and characterizations, electronics, optics, photonics, magnetics, magnonics, and their non-equilibrium physics, devices and applications, machine learning, and other advanced data science techniques.
Research Centers
-
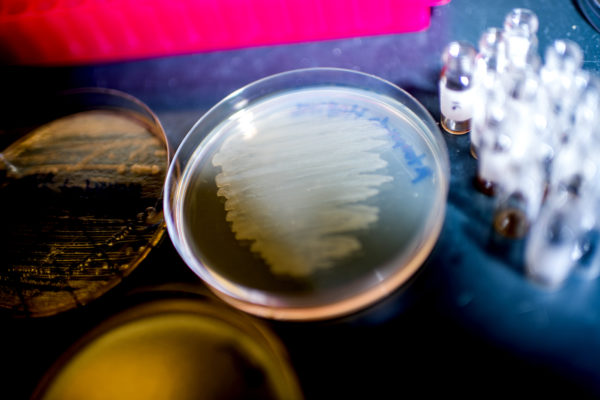
The Center translates basic discoveries into novel antimicrobial therapies to combat Biowarfare and conventional pathogen threats. The rise of multidrug resistant pathogens and the threat of genetically engineered bioweapons represent an urgent need for antimicrobial therapies. The Center is funded by grants from the NIH, NSF, and DOE.
-

The Brain Game Center for Mental Fitness and Well-Being designs interactive experiences for diverse populations to transform how we think about mental fitness and well-being. Our team conducts groundbreaking research, rigorous testing, and widespread distribution of scientifically refined software to help numerous people address their cognitive needs and reach their cognitive goals.
-

As a consortium of research institutions, healthcare systems and private companies, Epistorm develops innovative methodologies to integrate high-resolution mobility, airline travel, genomic and wastewater surveillance data—with mechanistic, statistical, and deep learning forecasting models to increase the accuracy of predictive analytics to help the US make more informed decisions during future outbreaks of infectious diseases.
-

The objective of the Center for Complex Network Research (CCNR) is simple: think networks. Research focuses on how networks emerge/evolve, how they look, and how they impact our understanding of complex systems. CCNR’s research has developed to unexpected areas, including the topology of the World Wide Web; complex networks inside the cell, and the Internet’s Achilles’ Heel.
-

The Center for Complex Particle Systems makes, measures, and models complex materials from molecular, nano, and microscale components combining order and disorder to establish simple chemical pathways to resource-conscious, multifunctional and intelligent materials for a sustainable and healthy future. By bridging the theory of graphs, physics of networks, science of particles, and examples of evolution, we create the foundations of their engineering and develop its diverse global practitioners.
-

The Center for Drug Discovery is dedicated to the discovery of novel medications and the development of approaches and technologies aimed at improving the discovery of new therapeutic drugs.
-

Center for Innovation in Mathematics, Physics, and Computational Theory
The Center for Innovation in Mathematics, Physics, and Computational Theory (IMPACT) advances interdisciplinary research at the intersections of theoretical physics, mathematics, and computational science. The center fosters bidirectional collaboration, applying AI and quantum information methods to fundamental problems in math and physics while bringing mathematical and physical techniques to bear on challenges in AI and quantum computing.
-

The Center for Interdisciplinary Research on Complex Systems fosters collaborations between researchers from different scientific and engineering disciplines who share a common interest in elucidating fundamental aspects of the structure and function of complex physical and biological systems across multiple levels of organization using a combination of quantitative state-of-the-art experimental and theoretical research tools.
-

The Center for Marine Ecological Genomics will develop solutions for studying genetic diversity in marine and coastal systems, and advance frameworks for translating
those solutions into practical applications for the sustainability of natural resources. By developing an open science ecosystem for marine genomics research and leveraging
advancements in AI, we will accelerate genomic discoveries from non-model marine organisms. -

The Center for Theoretical Biological Physics (CTBP) uses methods from physics, mathematics, and chemistry to understand biological problems. Research within CTBP focuses on broadly defined areas, including the fundamentals of physical genetics, interacting active elements and biological functionality, integrated living systems, and the development of core methodology.
-
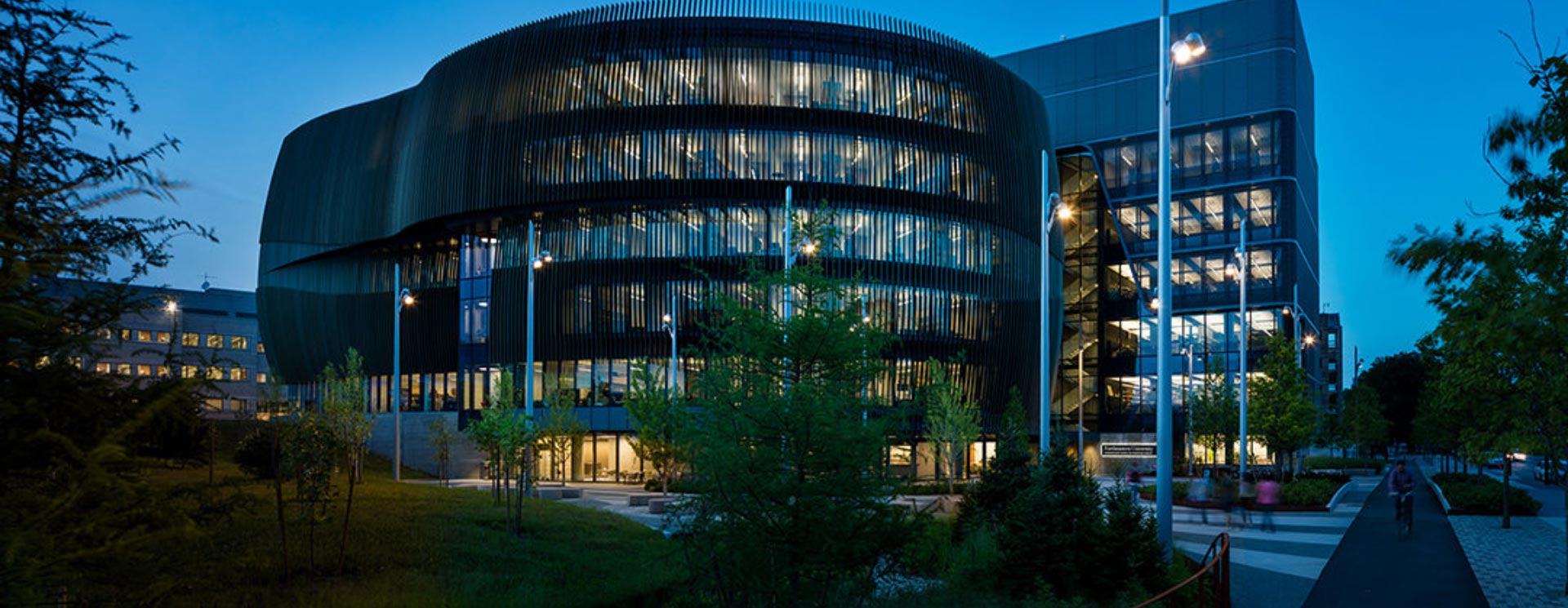

The Center for Translational NeuroImaging endeavors to provide services to the academic community interested in animal modeling and drug testing to aid in the diagnosis and treatment of CNS diseases. The center is also committed to training the next generation of imaging scientists to meet the needs of the pharmaceutical and biotechnology industries.
-

The Cross-College Magnetics Center will advance the frontiers of magnetic science and technology while fostering an interdisciplinary environment for collaboration and provide educational training for students at all levels. As a global hub for magnetics research, the Magnetics Center at Northeastern aims to accelerate scientific discovery and technological innovation in the field.
-
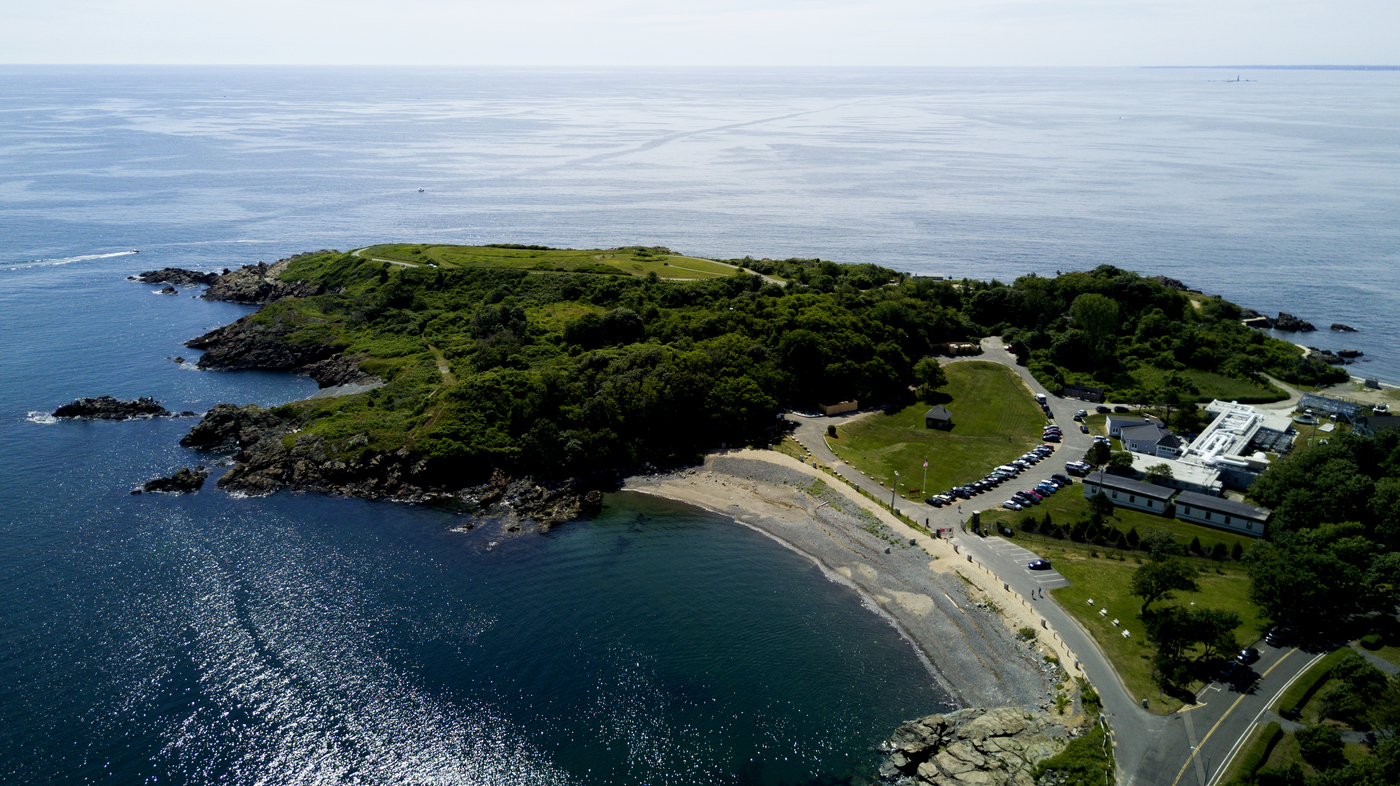

Located on the Nahant campus, the Marine Science Center is an internationally recognized research institution that focuses on the ocean environment, marine life and ecology, and discovering biotechnological and medical potentials in the sea. Projects include building underwater robots and creating genetically engineered seaweed to clean wastewater from agriculture facilities.
-

The Nanomedicine Innovation Centers provides a collaborative, innovative environment with state-of-the-art facilities for materials education and cutting-edge research in nanomedicine.
-
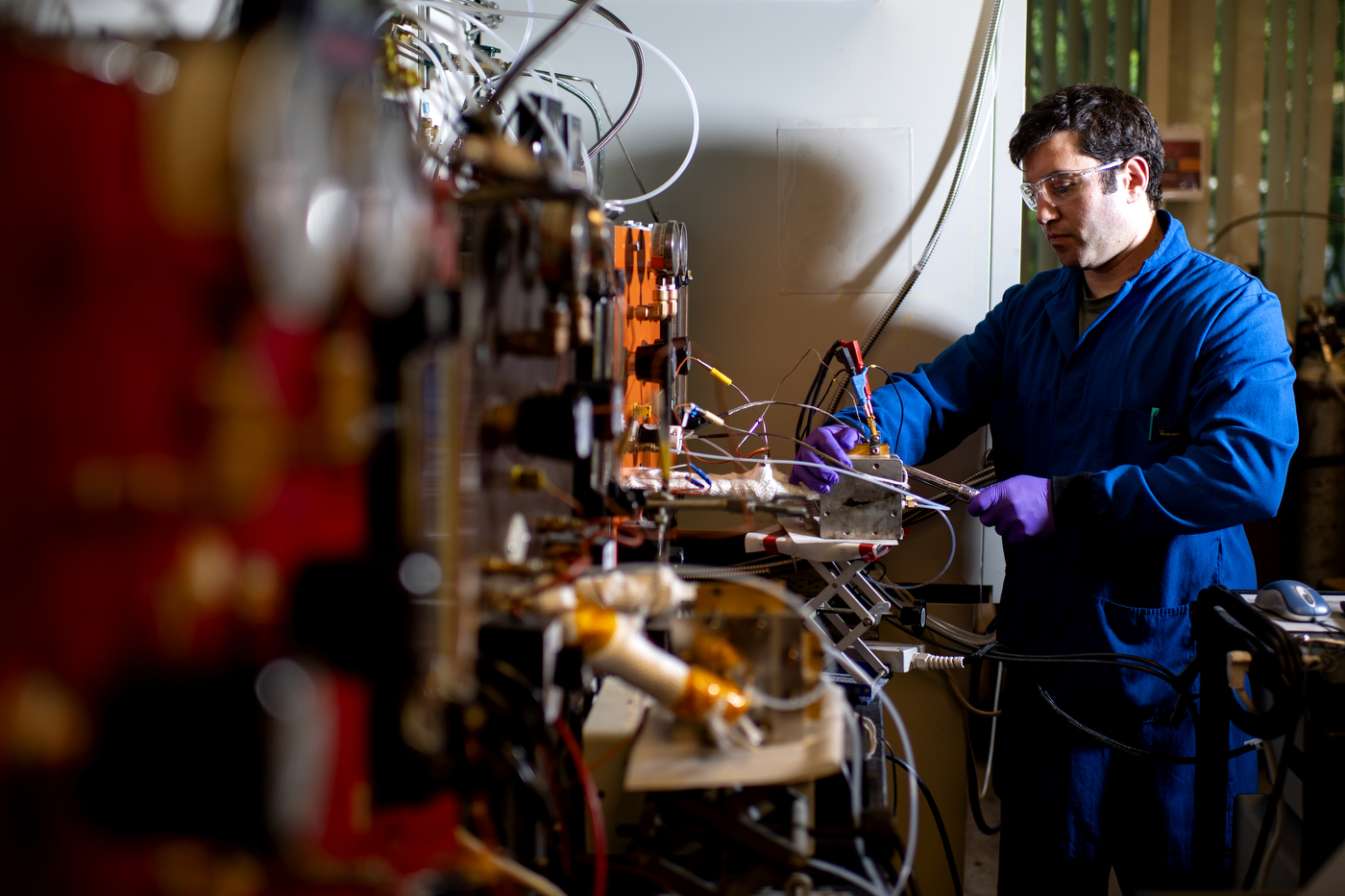

The Northeastern University Center for Renewable Energy Technology aims to be at the frontier of science and technology of clean energy conversion and storage. The range of efforts includes materials science, advanced in situ spectroscopy, micro-fabrication methods and manufacturing technology.
-
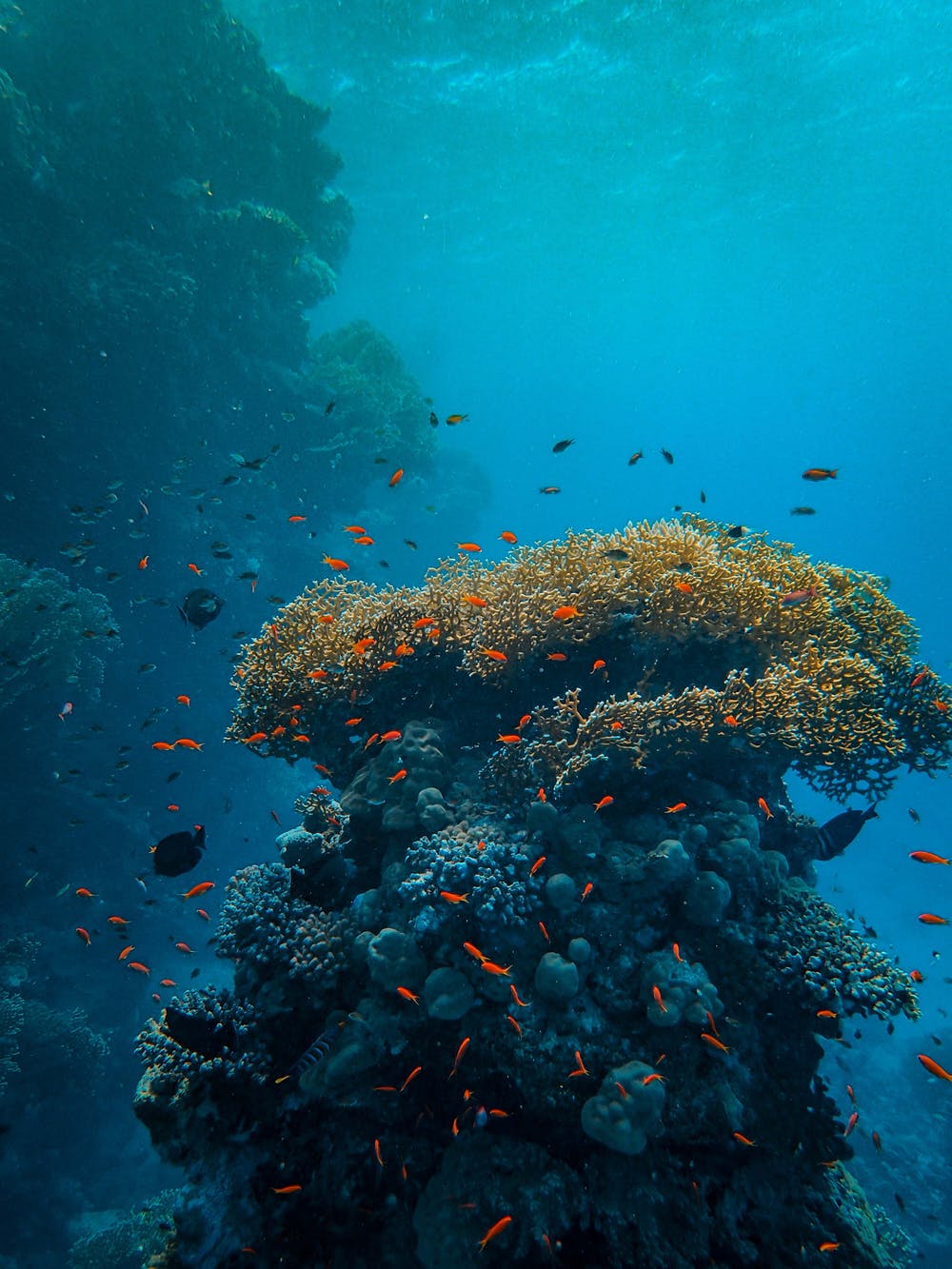

The Ocean Genome Legacy Center at Northeastern University’s Marine Science Center is a non-profit research organization and biological specimen repository dedicated to exploring and preserving the wealth of information contained in the genomes of endangered, rare, unusual, and ecologically critical marine organisms. Its mission is to acquire, authenticate, study, preserve, develop, and distribute genetic materials, biological specimens, and information to the research community.
-

The Plastics Center promotes responsible and sustainable plastic use through innovative research and collaboration to address global needs. It will drive solutions through research, policy, and education, connecting science, engineering, arts, and policy. The center aims to develop safer practices, reduce pollution, and empower communities, collaborating with academia, business, and government to create sustainable solutions and advance health.
Explore Northeastern’s Center for STEM Education
Departments
Research Consortia
-
Healthy Aging
Our diverse group of researchers with expertise in aging answer important questions about the well-being of the global aging population by combining personalized molecular genetics, biobehavioral sciences, computational modeling, health data analytics, bioengineered robotic devices, and lifestyle interventions that collectively target “health-span”.
-

Mathematical Modeling
Mathematical models can bridge the gap between theory and empirical data with far-reaching implications, solving complex problems across disciplines. Through their diverse expertise, Northeastern faculty tackle a wide array of challenges, from understanding the dynamics of biological systems to modeling the behavior of materials at the nanoscale.
-

Mechanobiology
Comprising interdisciplinary experts from fields such as biology, engineering, physics, and materials science, Northeastern researchers delve into the fundamental mechanisms by which mechanical forces influence and regulate biological processes at various scales.
-

Understanding and improving mental health is a multi-scale, multi-partner endeavor. By engaging with interdisciplinary research at all levels – across departments, colleges, institutions, cities, and regions – we are tackling some of the most important societal challenges.
-
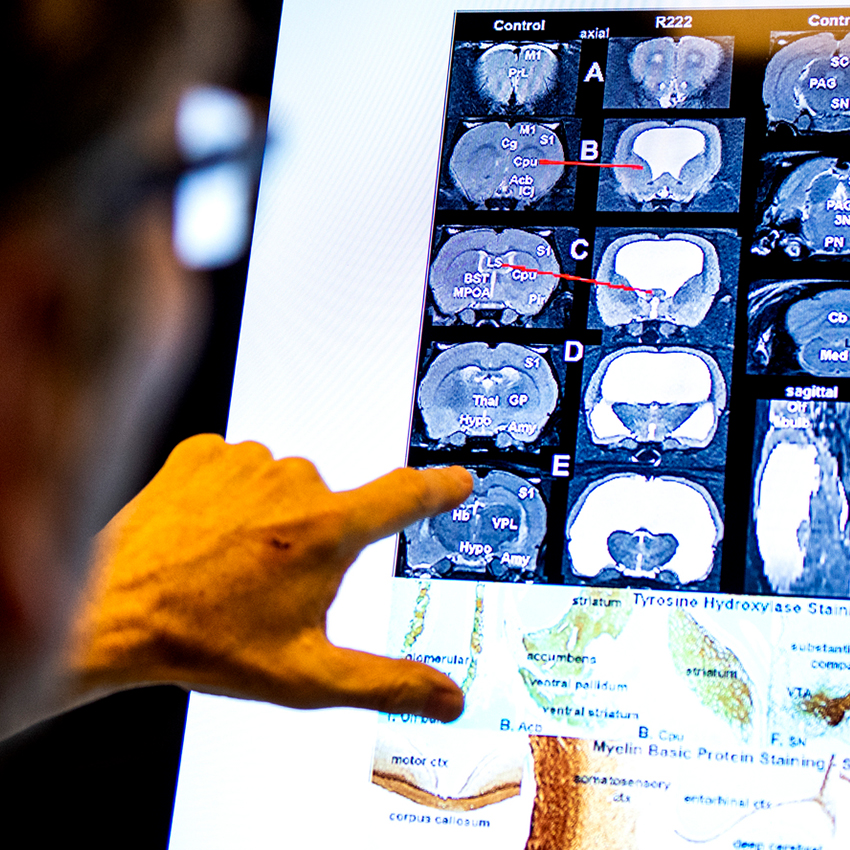
Metabolism
Utilizing cutting-edge techniques, including neuroimaging and metabolic profiling, our researchers aim to decipher the complex interplay between metabolism, brain function, and mental health outcomes. Their research spans a wide range of critical areas, from investigating the impact of nutrition on mood and cognitive function to uncovering the role of metabolic dysregulation in psychiatric disorders such as depression, anxiety, and schizophrenia
-

Quantum
Quantum research spans a wide spectrum of phenomena, from quantum computing and sensing to the development of novel quantum materials. Utilizing cutting-edge experimental setups and theoretical and computational models, we are investigating the practical applications of quantum principles and their potential to revolutionize material science, medicine, and our understanding of the universe.
Shared User Facilities
-

The Biomedical Imaging Center is a state-of-the-art magnetic resonance imaging (MRI) facility with cutting-edge acquisition and analysis techniques to support biomedical imaging research projects that use MRI.
-

The Boston Electron Microscopy Center maintains three electron microscopes including high-resolution field emission scanning electron microscope Hitachi S-4800 FE-SEM, high-resolution transmission electron microscope JEOL 2010F TEM and high-contrast JEOL JEM-1010 TEM as well as a biological sample preparation lab.
-
Researchers at the institute are pioneering new imaging technologies to reveal, in real time, chemical processes in the brain and body, leading to a deeper understanding of how diseases unfold and faster, more precise diagnoses and treatments.
-

The Center for Translational NeuroImaging endeavors to provide services to the academic community interested in animal modeling and drug testing to aid in the diagnosis and treatment of CNS diseases.
-

The Experiential Quantum Advancement Laboratories (EQUAL) is a surface science lab equipped to design, synthesize, and discover spin-resolved phenomena in real and reciprocal space using state-of-the-art connected instruments.
-
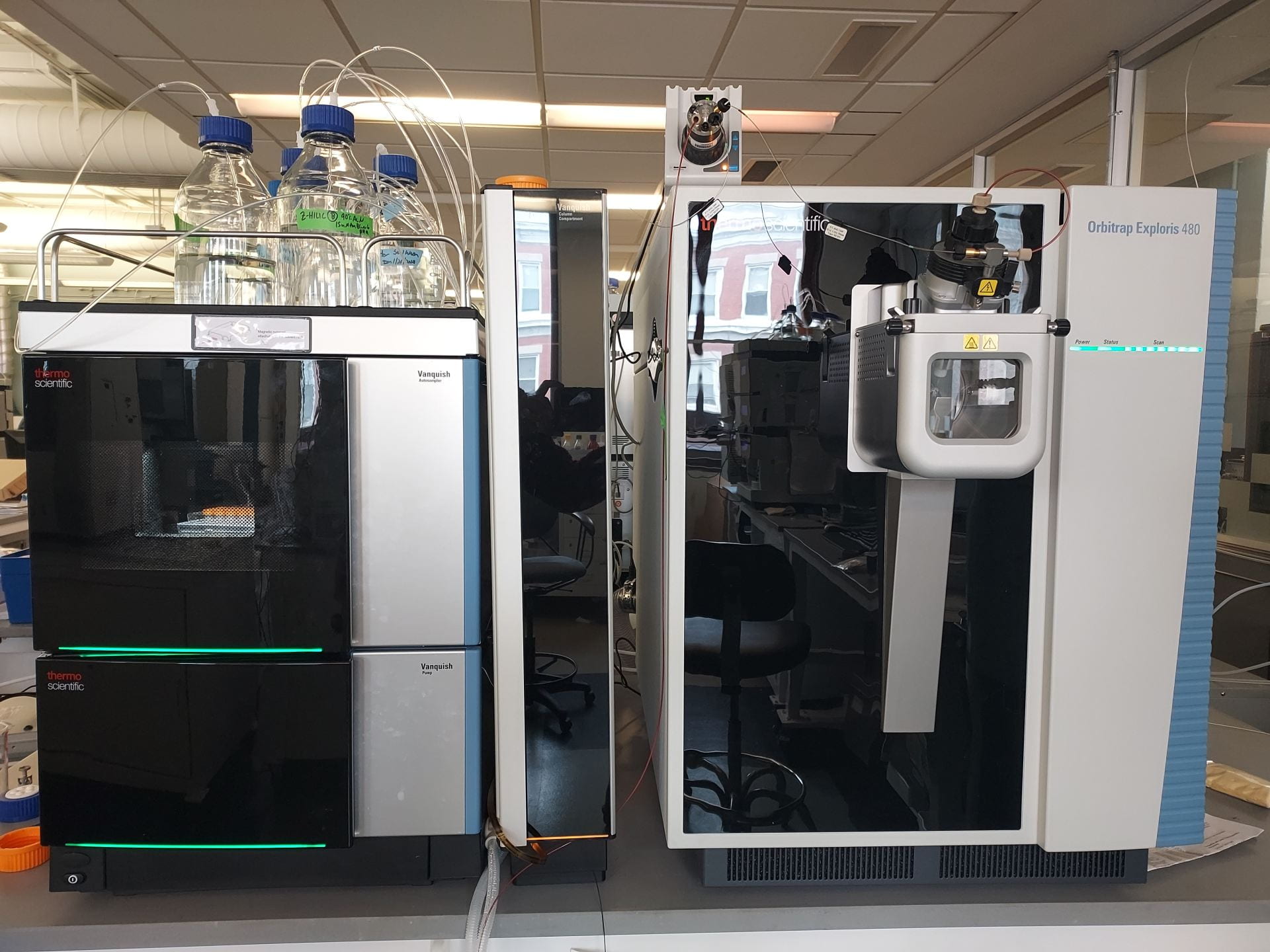

The Mass Spectrometry Facility has three Thermo Orbitraps dedicated to proteomics, small molecule analysis and native protein analysis and one Thermo triple quadrupole for targeted analysis of proteins or small molecules.
-

The Nuclear Magnetic Resonance Facility maintains five NMR spectrometers and one EPR spectrometer for teaching and sponsored research.
-

Equipment at the Wet Lab Makerspace include a Flow Cytometer, Inverted Fluorescent Microscope, Multimeter for pH, Conductivity, and DO, UV/Vis Spectrophotometer, and more to support projects in Cell Culture, Chemistry, Microscopy, and Molecular Biology.
Be a part of the next era of discovery
Conduct postdoctoral research in the College of Science.