Northeastern University’s Mass Spectrometry Facility has been awarded a $2.2 million research infrastructure grant from the Massachusetts Life Sciences Center (MLSC) to build a cutting-edge core supporting single-cell proteomics research. The award, through the Center’s Research Infrastructure program, will establish a fully integrated pipeline that enables researchers to measure proteins at single-cell resolution, filling a critical gap in cellular biology research.
Under the direction of Drs. Virginie Sjoelund and Alexander Ivanov (Co-PIs of this award), the facility will deploy state-of-the-art instrumentation that addresses longstanding technical challenges in the field and positions Northeastern as a regional hub for advanced proteomics capabilities.
Bridging the Protein Gap in Single-Cell Biology
While single-cell transcriptomics has advanced rapidly in recent years, the field has faced a major limitation: proteins, which carry out cellular functions, have been extremely difficult to measure at single-cell resolution. This gap has prevented researchers from better understanding cellular heterogeneity, rare cell populations, and spatial organization in tissues.
“At Northeastern and other research institutions, many researchers wanted to explore questions that couldn’t be answered at the single-cell level with RNA alone, but they didn’t have access to the necessary instrumentation or expertise,” explained Dr. Sjoelund.
The new centralized single-cell proteomics research facility will integrate precise cell isolation, automated sample preparation, and highly sensitive mass spectrometry to generate protein-level data that complement transcriptomics and open new opportunities to study functional states, signaling pathways, and tissue organization.
Minimizing Loss, Maximizing Data Quality
Single-cell proteomics presents extraordinary technical demands. Each cell contains only minute amounts of protein, and even small losses during processing can compromise data quality. Achieving reproducible, quantitative measurements while integrating spatial information from tissues has been a major hurdle for the field.
The new infrastructure addresses these challenges through a comprehensive approach. Highly sensitive mass spectrometry, combined with precise single-cell isolation and automated low-volume sample preparation, work together to minimize sample loss and maximize reproducibility, making high-quality single-cell and spatial proteomics achievable for a wide range of researchers at Northeastern and nationwide.
A Complete Proteomics Pipeline
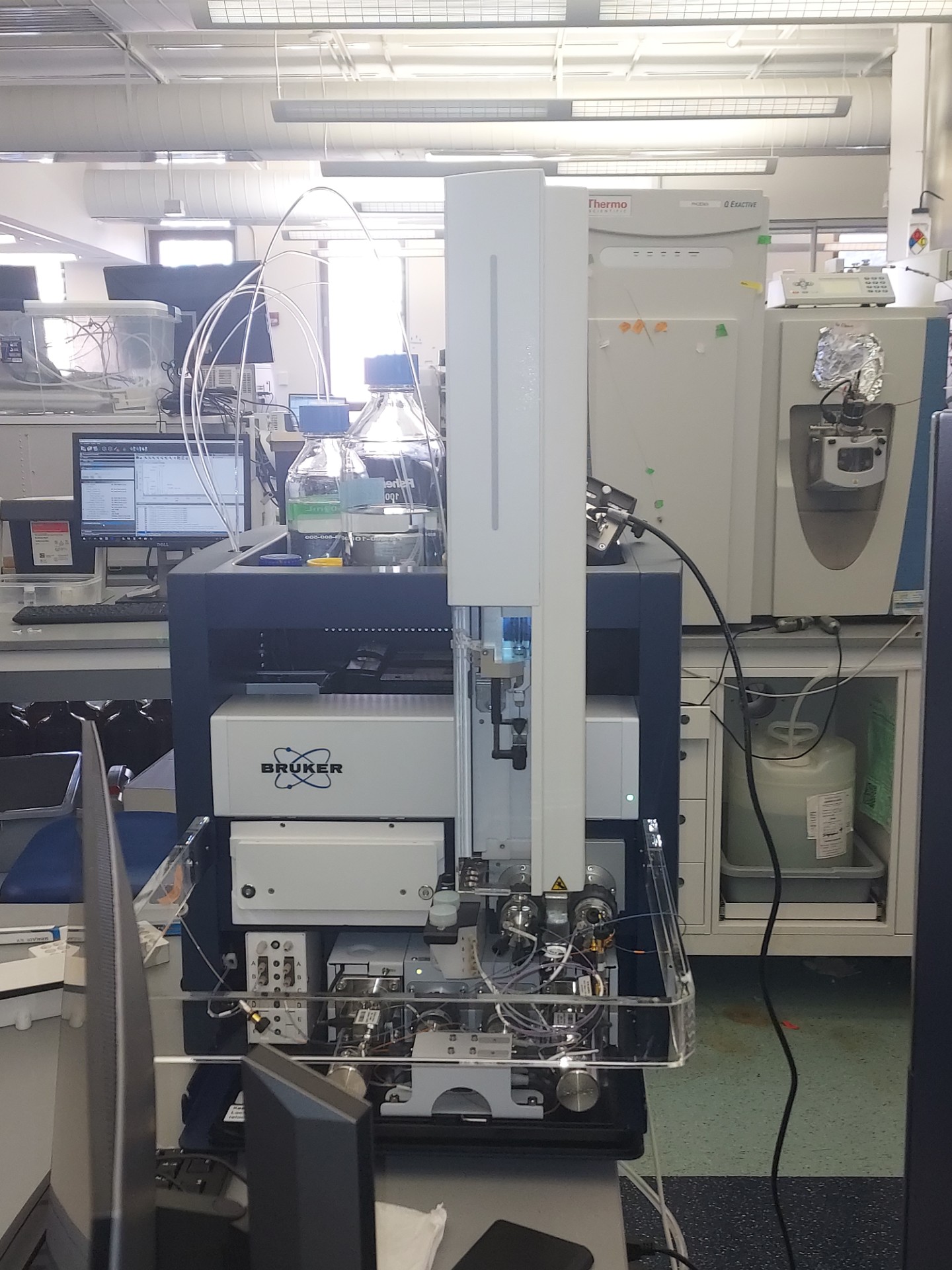
The awarded MLSC research infrastructure grant supports four key instruments that create a seamless workflow from cell selection to protein measurement.
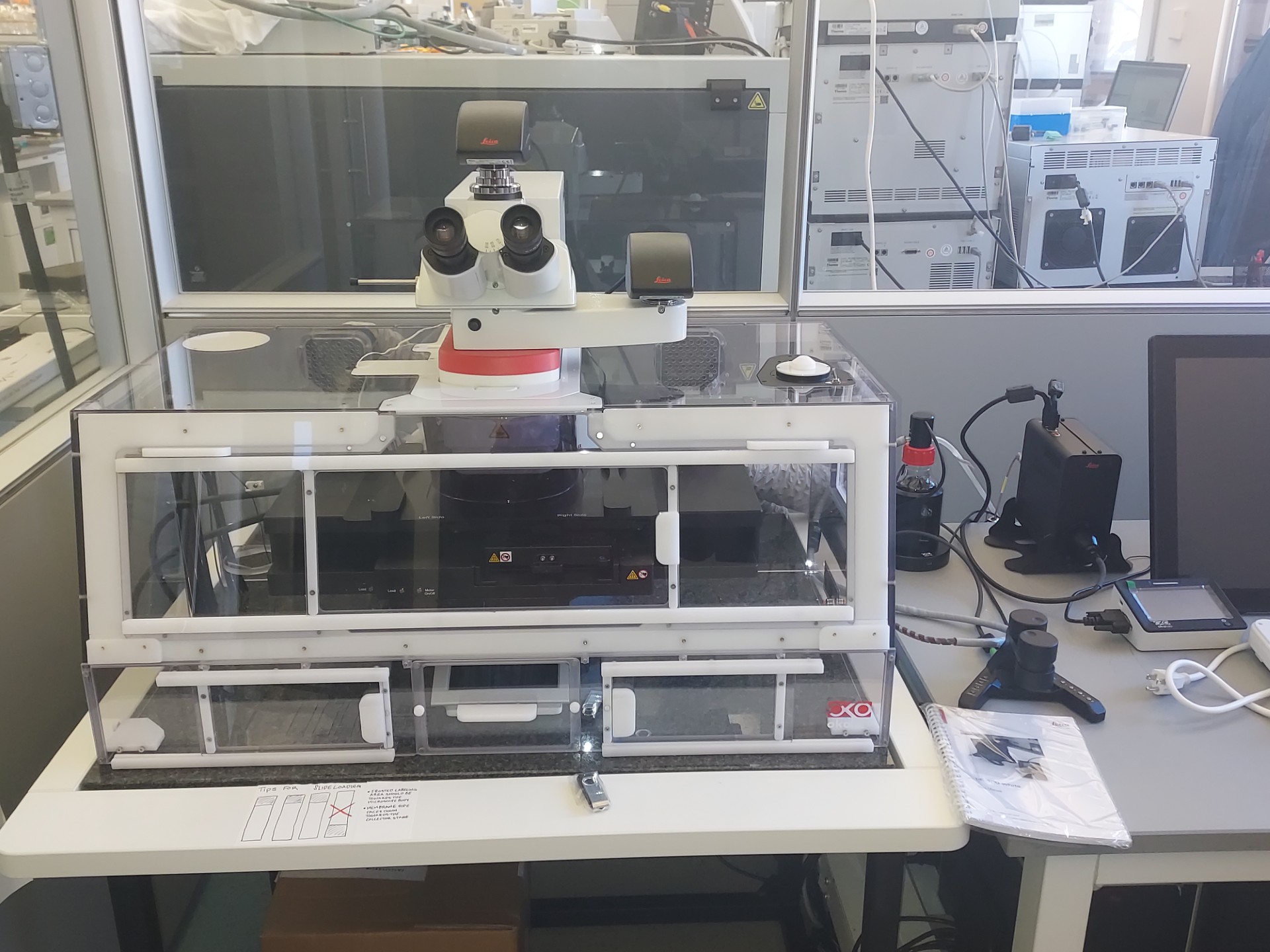
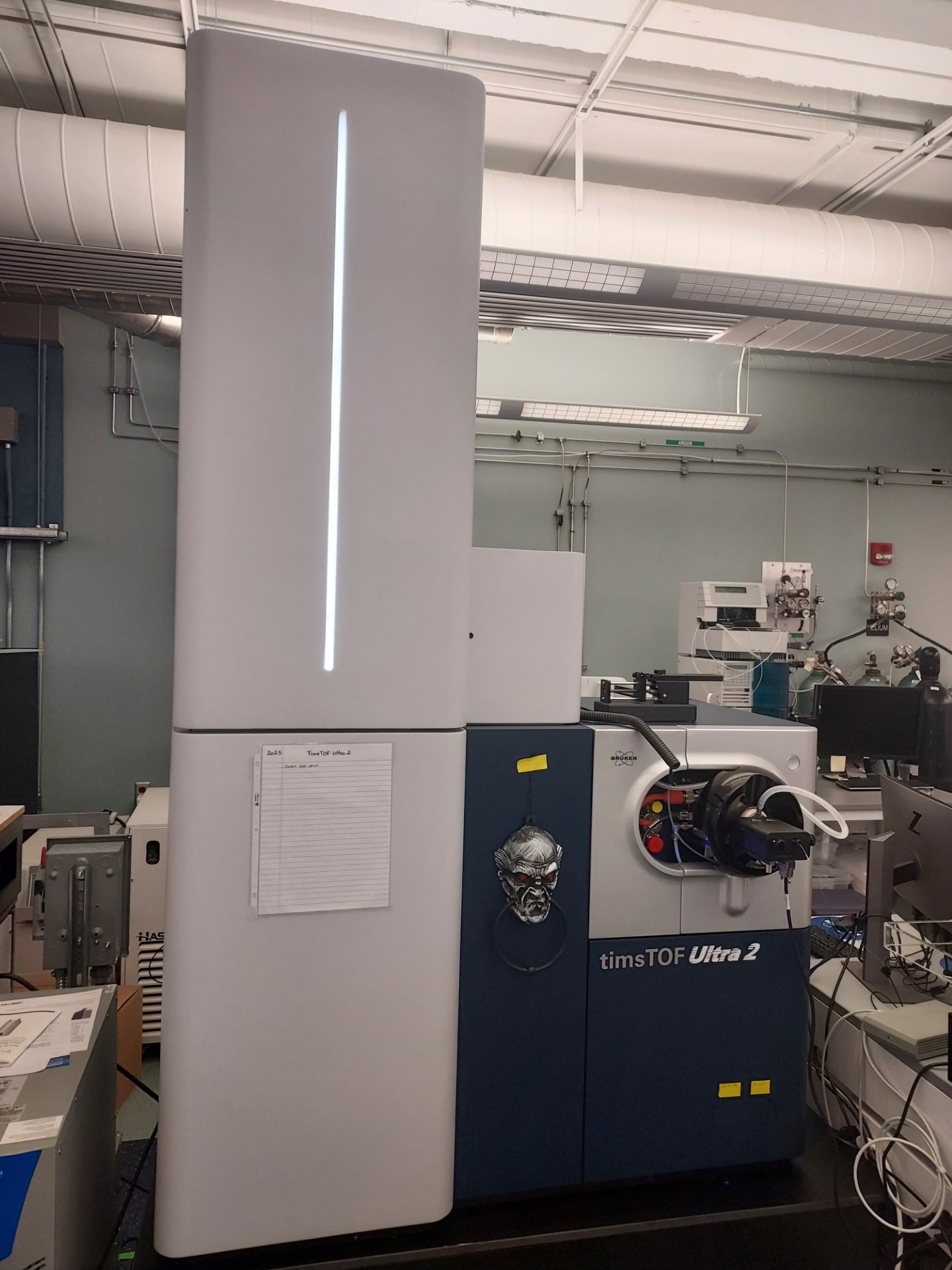
A Bruker TimsTOF Ultra 2 mass spectrometer provides highly sensitive protein quantification, while a Leica LMD7 laser microdissection microscope isolates cells or tissue regions with spatial context. A Scienion CellenONE enables precise and automated single-cell isolation and processing, and a Tecan UNO handles nanoliter-scale automated sample preparation.
Together, these instruments enable experiments that were previously extremely difficult or impossible.

Building a Regional Resource
Over the next three to five years, the facility aims to establish standardized, reproducible workflows for single-cell and spatial proteomics and make them accessible to the broader research community. The facility will support diverse projects ranging from fundamental biology research to translational studies and provide training to help investigators, who are new to these methods, get up to speed in their single-cell experiments.
“We want to integrate proteomics with other single-cell technologies, including imaging and orthogonal molecular profiling approaches, and position the facility as a regional resource that fosters collaborations and attracts additional funding,” said Dr. Sjoelund.
“We believe this will be the first shared research facility at an academic institution offering single-cell proteomics analyses in the state of Massachusetts and across the US, and we are excited to be trailblazers in this new and rapidly developing area of research,” said Prof. Ivanov.
Opening Doors to Single-Cell Discovery
Researchers will access the facility through a core-facility model, with consultation on experimental design, fee-for-service access, and assisted or full-service options for complex experiments. Training workshops and hands-on sessions will be available for those who want to learn the workflows, with the goal of making these capabilities available to both internal investigators and external collaborators.
The methods are particularly well-suited for projects where protein function, signaling, or spatial context is critical, such as studies of tumor heterogeneity, immune cell states, or rare cell populations in development or disease. They are also valuable for investigating post-translational modifications and dynamic cell states that cannot be inferred from RNA alone.
Pilot Projects to Showcase Potential
To showcase the facility’s potential, the team plans to initiate collaborative pilot projects with selected investigators. These pilot studies will help demonstrate feasibility, generate high-quality proof-of-concept data, and refine protocols for different sample types. They will also serve as examples for the broader research community, encouraging wider adoption of single-cell proteomics approaches.
The research infrastructure investment by the MLSC marks a significant milestone in making advanced proteomics capabilities accessible to researchers across the region, enabling new discoveries in cellular biology, disease mechanisms, and therapeutic development.
“This award is a testament to the innovative work in our department and was led by Prof. Ivanov and Dr. Sjoelund to bring unique capabilities to Northeastern and the entire region,” remarks Dr. Penny Beuning, professor and chair of chemistry and chemical biology. “Single-cell proteomics is at the intersection of chemistry, biology, and data science. This facility will enable our faculty and collaborators to use multidisciplinary approaches to take on fundamental questions about cell function, with implications for health and disease.”